Natural language generation has improved remarkably over the last decade and OpenAI’s ChatGPT showcases an unbelievable performance of state-of-the art systems. Like many curious people over the last month, I played with ChatGPT from time to time. It’s super impressive – it can create poems, write prose in nearly any style, and provide detailed answers to all sorts of questions. Oh, and it’s also an amazing bullshit artist and it will lie with abandon.
I decided to ask ChatGPT a specific question about particle physics that is a bit subtle but has a straightforward answer. ChatGPT’s response is stunning bullshit.

ChatGPT gets the first part right – the most common decay mode is to an electron, an electron antineutrino, and a muon neutrino. Now, the rest of the answer is completely wrong. First, the muon is kinematically forbidden from decaying into pions. The muon is simply not massive enough to do so. Second, even if the muon were massive enough to decay with three charged pions in the final state (like the tau lepton can), conservation of lepton number still requires neutrinos. Apparently ChatGPT has thrown lepton number overboard so that it can give me an answer!
The correct answer to my quetsion is: Theoretically yes, but hopelessly rare. No observation of muon decay without emitting neutrinos has ever been observed.
In the Standard Model (the theory of fundamental particle interactions) with massless neutrinos, muon decay must include neutrinos in the final state. The Standard Model with massless neutrinos conserves family lepton number and so neutrinos in the final state are required to conserve muon and electron number (a muon neutrino and an electron antineutrino). However, neutrino flavour oscillation observations tell us that neutrinos have very, very tiny masses. Extending the Standard Model to include neutrino masses allows for the muon to decay without neutrinos (for example to an electron and a photon) but at rates so extremely small that we have no hope of observing neutrinoless decays of the muon unless the Standard Model is further extended with exotic physics.
Ok, rather technical stuff. But where did ChatGPT come up with the idea of pions in the final state? Well, I asked it. And sure enough ChatGPT came up with references, including a paper in Nature from 1982 with claims of actually observing neutrinoless decays of the muon! Well, either I missed something during my PhD and postdocs (I have done a fair amount of work on supersymmetric extensions of the Standard Model with enhanced flavour violating decays of the muon) or ChatGPT is making stuff up. All of the references that ChaptGPT gave me are complete fabrications – they don’t exist!!

It turns out that natural language generation in systems like ChatGPT suffer from hallucinating unintended text. Its creative power, its ability to write poems etc., requires such an enormous amount of flexibility that ChatGPT seems to start making things up to complete an answer. I’m not an expert on natural language processing, but I know that researchers are aware of the phenomenon.
I happen to know something about particle physics so it’s not a big deal that ChatGPT fibbed to answer my question. In fairness, my question is a bit subtle (short answer is No with an if, long answer is Yes with a but). But imagine circumstances that are more important. How do you know if you can trust the answers? Asking it for reference doesn’t seem to help.
To me the lesson is that systems like ChatGPT are going to be ever more useful with the potential to make life much more convenient. But these systems are not a substitute for human cognition. For the teachers out there – don’t worry about ChatGPT writing your students’ essays, just check to see if the references exist!
Update (February 23, 2023)
I returned to ChatGPT last night, getting into a discussion with it about the top quark. ChatGPT continued to give insane answers about QCD, electroweak physics, and the nature of the Higgs sector. I thought I could get it on track if it would just realize that the muon’s mass is less than the pions. It produced this gem:

ChatGPT fails the Piaget test.














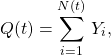
![Rendered by QuickLaTeX.com \begin{align*}\phi_{Q(t)}(u) &=\mathbb{E}[e^{iuQ(t)}] \\ &= \exp\left(rt\ln\left(\frac{1-p}{1-pe^{iu}}\right)\right) \\ &= \left(\frac{1-p}{1-pe^{iu}}\right)^{rt},\end{align*}](https://maybury.ca/the-reformed-physicist/wp-content/ql-cache/quicklatex.com-493e9b3cc164dbdfe835dadda287c3ca_l3.png)
![Rendered by QuickLaTeX.com \begin{align*}\mathbb{E}[e^{iuZ(t)}] &= \exp(-t\psi(-\eta(u))) \\ &= \left(1- \frac{r}{b}\ln\left(\frac{1-p}{1-pe^{iu}}\right)\right)^{-at},\end{align*}](https://maybury.ca/the-reformed-physicist/wp-content/ql-cache/quicklatex.com-b8dec93aa84eef4c5279cb4cf8a9011b_l3.png)
![Rendered by QuickLaTeX.com \begin{align*}\mathbb{E}[e^{iuQ(t)}] &= \exp(t\,\eta(u)) \\\mathbb{E}[e^{-sT(t)}] &= \exp(-t\,\psi(s)), \end{align*}](https://maybury.ca/the-reformed-physicist/wp-content/ql-cache/quicklatex.com-a388ec11ca6d71673b206a186378daf0_l3.png)