Last month my collaborators, Jerome Levesque and David Shaw, and I built a branching process model for describing Covid-19 propagation in communities. In my previous blog post, I gave a heuristic description of how the model works. In this post, I want to expand on some of the technical aspects of the model and show how the model can be calibrated to data.
The basic idea behind the model is that an infected person creates new infections throughout her communicable period. That is, an infected person “branches” as she generates “offspring”. This model is an approximation of how a communicable disease like Covid-19 spreads. In our model, we assume that we have an infinite reservoir of susceptible people for the virus to infect. In reality, the susceptible population declines – over periods of time that are much longer than the communicable period, the recovered population pushes back on the infection process as herd immunity builds. SIR and other compartmental models capture this effect. But over the short term, and especially when an outbreak first starts, disease propagation really does look like a branching process. The nice thing about branching processes is that they are stochastic, and have lots of amazing and deep relationships that allow you to connect observations back to the underlying mechanisms of propagation in an identifiable way.
In our model, both the number of new infections and the length of the communicable period are random. Given a communicable period, we model the number of infectious generated, ![]() , as a compound Poisson process,
, as a compound Poisson process,
(1) 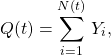
where ![]() is the number of infection events that arrived up to time
is the number of infection events that arrived up to time ![]() , and
, and ![]() is the number infected at each infection event. We model
is the number infected at each infection event. We model ![]() with the logarithmic distribution,
with the logarithmic distribution,
(2) ![]()
which has mean, ![]() . The infection events arrive exponentially distributed in time with arrival rate
. The infection events arrive exponentially distributed in time with arrival rate ![]() . The characteristic function for
. The characteristic function for ![]() reads,
reads,
(3) ![Rendered by QuickLaTeX.com \begin{align*}\phi_{Q(t)}(u) &=\mathbb{E}[e^{iuQ(t)}] \\ &= \exp\left(rt\ln\left(\frac{1-p}{1-pe^{iu}}\right)\right) \\ &= \left(\frac{1-p}{1-pe^{iu}}\right)^{rt},\end{align*}](https://maybury.ca/the-reformed-physicist/wp-content/ql-cache/quicklatex.com-493e9b3cc164dbdfe835dadda287c3ca_l3.png)
with ![]() and thus
and thus ![]() follows a negative binomial process,
follows a negative binomial process,
(4) ![]()
The negative binomial process is important here. Clinical observations suggest that Covid-19 is spread mostly by a minority of people in large quantities. Research suggests that the negative binomial distribution describes the number of infections from infected individuals. In our process, during a communicable period, ![]() , an infected individual infects
, an infected individual infects ![]() people based on a draw from the negative binomial with mean
people based on a draw from the negative binomial with mean ![]() . The infection events occur continuously in time according to the Poisson arrivals. However, the communicable period,
. The infection events occur continuously in time according to the Poisson arrivals. However, the communicable period, ![]() , is in actuality a random variable,
, is in actuality a random variable, ![]() , which we model as a gamma process,
, which we model as a gamma process,
(5) ![]()
which has a mean of ![]() . By promoting the communicable period to a random variable, the negative binomial process changes into a Levy process with characteristic function,
. By promoting the communicable period to a random variable, the negative binomial process changes into a Levy process with characteristic function,
(6) ![Rendered by QuickLaTeX.com \begin{align*}\mathbb{E}[e^{iuZ(t)}] &= \exp(-t\psi(-\eta(u))) \\ &= \left(1- \frac{r}{b}\ln\left(\frac{1-p}{1-pe^{iu}}\right)\right)^{-at},\end{align*}](https://maybury.ca/the-reformed-physicist/wp-content/ql-cache/quicklatex.com-b8dec93aa84eef4c5279cb4cf8a9011b_l3.png)
where ![]() , the Levy symbol, and
, the Levy symbol, and ![]() , the Laplace exponent, are respectively given by,
, the Laplace exponent, are respectively given by,
(7) ![Rendered by QuickLaTeX.com \begin{align*}\mathbb{E}[e^{iuQ(t)}] &= \exp(t\,\eta(u)) \\\mathbb{E}[e^{-sT(t)}] &= \exp(-t\,\psi(s)), \end{align*}](https://maybury.ca/the-reformed-physicist/wp-content/ql-cache/quicklatex.com-a388ec11ca6d71673b206a186378daf0_l3.png)
and so,
(8) 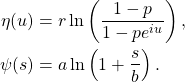
![]() is the random number of people infected by a single infected individual over her random communicable period and is further over-dispersed relative to a pure negative binomial process, getting us closer to driving propagation through super-spreader events . The characteristic function in eq.(6) for the number of infections from a single infected person gives us the entire model. The basic reproduction number
is the random number of people infected by a single infected individual over her random communicable period and is further over-dispersed relative to a pure negative binomial process, getting us closer to driving propagation through super-spreader events . The characteristic function in eq.(6) for the number of infections from a single infected person gives us the entire model. The basic reproduction number ![]() is,
is,
(9) ![]()
From the characteristic function we can compute the total number of infections in the population through renewal theory. Given a random characteristic ![]() , such as the number of infectious individuals at time
, such as the number of infectious individuals at time ![]() , (e.g.,
, (e.g., ![]() where
where ![]() is the random communicable period) the expectation of the process follows,
is the random communicable period) the expectation of the process follows,
(10) ![]()
where ![]() is the counting process (see our paper for details). When an outbreak is underway, the asymptotic behaviour for the expected number of counts is,
is the counting process (see our paper for details). When an outbreak is underway, the asymptotic behaviour for the expected number of counts is,
(11) ![]()
where,
(12) 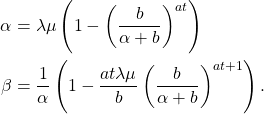
The parameter ![]() is called the Malthusian parameter and it controls the exponential growth of the process. Because the renewal equation gives us eq.(11), we can build a Bayesian hierarchical model for inference with just cumulative count data. We take US county data, curated by the New York Times, to give us an estimate of the Malthusian parameter and therefore the local
is called the Malthusian parameter and it controls the exponential growth of the process. Because the renewal equation gives us eq.(11), we can build a Bayesian hierarchical model for inference with just cumulative count data. We take US county data, curated by the New York Times, to give us an estimate of the Malthusian parameter and therefore the local ![]() -effective across the United States. We use clinical data to set the parameters of the gamma distribution that controls the communicable period. We treat the counties as random effects and estimate the model using Gibbs sampling in JAGS. Our estimation model is,
-effective across the United States. We use clinical data to set the parameters of the gamma distribution that controls the communicable period. We treat the counties as random effects and estimate the model using Gibbs sampling in JAGS. Our estimation model is,
(13) 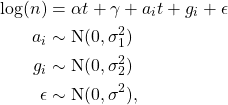
where ![]() is the county label; the variance parameters use half-Cauchy priors and the fixed and random effects use normal priors. We estimate the model and generate posterior distributions for all parameters. The result for the United States using data over the summer is the figure below:
is the county label; the variance parameters use half-Cauchy priors and the fixed and random effects use normal priors. We estimate the model and generate posterior distributions for all parameters. The result for the United States using data over the summer is the figure below:

Over the mid-summer, we see that the geographical distribution of ![]() across the US singles out the Midwestern states and Hawaii as hot-spots while Arizona sees no county with exponential growth. We have the beginnings of a US county based app which we hope to extend to other countries around the world. Unfortunately, count data on its own does not allow us to resolve the parameters of the compound Poisson process separately.
across the US singles out the Midwestern states and Hawaii as hot-spots while Arizona sees no county with exponential growth. We have the beginnings of a US county based app which we hope to extend to other countries around the world. Unfortunately, count data on its own does not allow us to resolve the parameters of the compound Poisson process separately.
If we have complete information, which might be possible in a small community setting, like northern communities in Canada, prisons, schools, or offices, we can build a Gibbs sampler to estimate all the model parameters from data without having to rely on the asymptotic solution of the renewal equation.
Define a complete history of an outbreak as a set of ![]() observations taking the form of a 6-tuple:
observations taking the form of a 6-tuple:
(14) ![]()
where,
![]() : index of individual,
: index of individual, ![]() : index of parent,
: index of parent, ![]() : time of birth,
: time of birth, ![]() : time of death,
: time of death, ![]() : number of offspring birth events,
: number of offspring birth events, ![]() : number of offspring.
: number of offspring.
With the following summary statistics:
(15) 
we can build a Gibbs sampler over the models parameters as follows:
(16) 
where ![]() are hyper-parameters.
are hyper-parameters.
Over periods of time that are comparable to the communicable window, such that increasing herd immunity effects are negligible, a pure branching process can describe the propagation of Covid-19. We have built a model that matches the features of this disease – high variance in infection counts from infected individuals with a random communicable period. We see promise in our model’s application to small population settings as an outbreak gets started.

Really interesting work! Can I ask a question? Was there a specific need for a Bayesian framework versus frequentist? A philosophical choice or something else? Just curious. Sometimes it’s not so clear to me why one approach is taken over another. Cheers,
Matt
Thanks, Matt. We rely heavily on mixed modelling techniques and Bayesian methods work rather naturally in those settings. There are ways to do mixed modelling without Bayesian methods – we use whatever takes best advantage of the data. If you would like to learn more about mixed modelling, including MCMC approaches, you might find Data Analysis Using Regression and Multilevel/Hierarchical Models, by Andrew Gelman, and Jennifer Hill worth a peek.